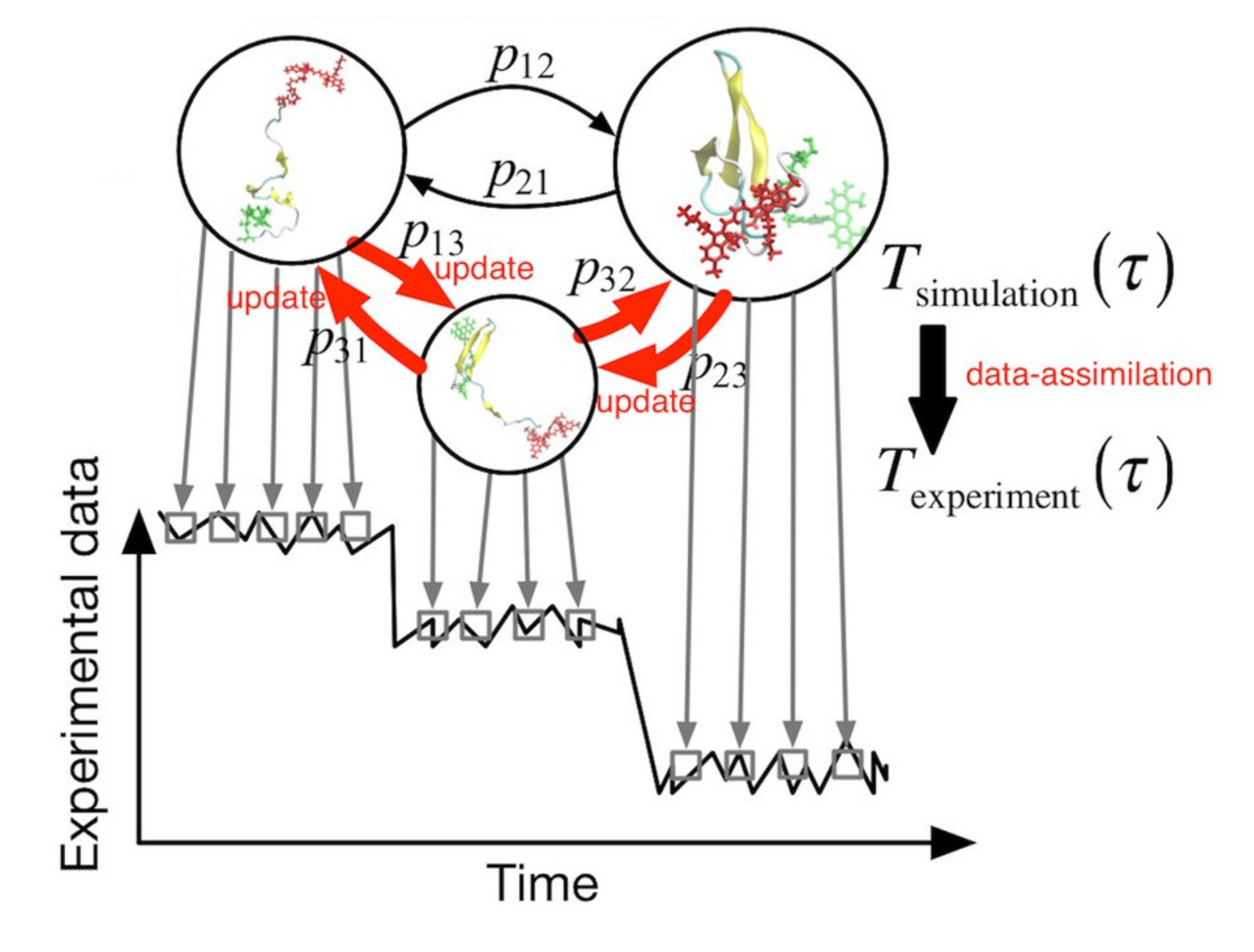
Many proteins involved in biological function go through large-scale structural changes for performing their role. We aim to unravel the mechanism of dynamic processes (dynamics) including structural changes in proteins. For this purpose, we developed a method for capturing processes involving various molecules such as proteins and solvent molecules, and implemented it into GENESIS. Using these methods, we elucidate the relation between protein structural dynamics and function. Moreover, we work on developing a new machine learning method connecting MD simulations and single molecule measurements, aiming for dynamics analysis which better reproduces experimental data.
References
- “Free Energy Analysis of a Conformational Change of Heme ABC Transporter BhuUV-T.”, Koichi Tamura and Yuji Sugita, J. Phys. Chem. Lett., 11, 2824-2829 (2020).
- “Chemo-mechanical Coupling in the Transport Cycle of a Type II ABC Transporter”, Koichi Tamura, Hiroshi Sugimoto, Yoshitsugu Shiro, Yuji Sugita, J. Phys. Chem. B, 123, 7270-7281 (2019).
- “Flexible selection of the solute region in replica exchange with solute tempering: Application to protein-folding simulations”, Motoshi Kamiya and Yuji Sugita., J. Chem. Phys., 149, 072304 (2018).
- “Linking time-series of single-molecule experiments with molecular dynamics simulations by machine learning”, Yasuhiro Matsunaga and Yuji Sugita, eLife, 7, e32668 (2018).
- “Refining Markov state models for conformational dynamics using ensemble-averaged data and time-series trajectories”, Yasuhiro Matsunaga and Yuji Sugita, J. Chem. Phys., 148, 241731 (2018).
- “Dimensionality of Collective Variables for Describing Conformational Changes of a Multi-Domain Protein”, Yasuhiro Matsunaga, Yasuaki Komuro, Chigusa Kobayashi, Jaewoon Jung, Takaharu Mori, and Yuji Sugita, J. Phys. Chem. Lett., 7, 1446–1451 (2016).